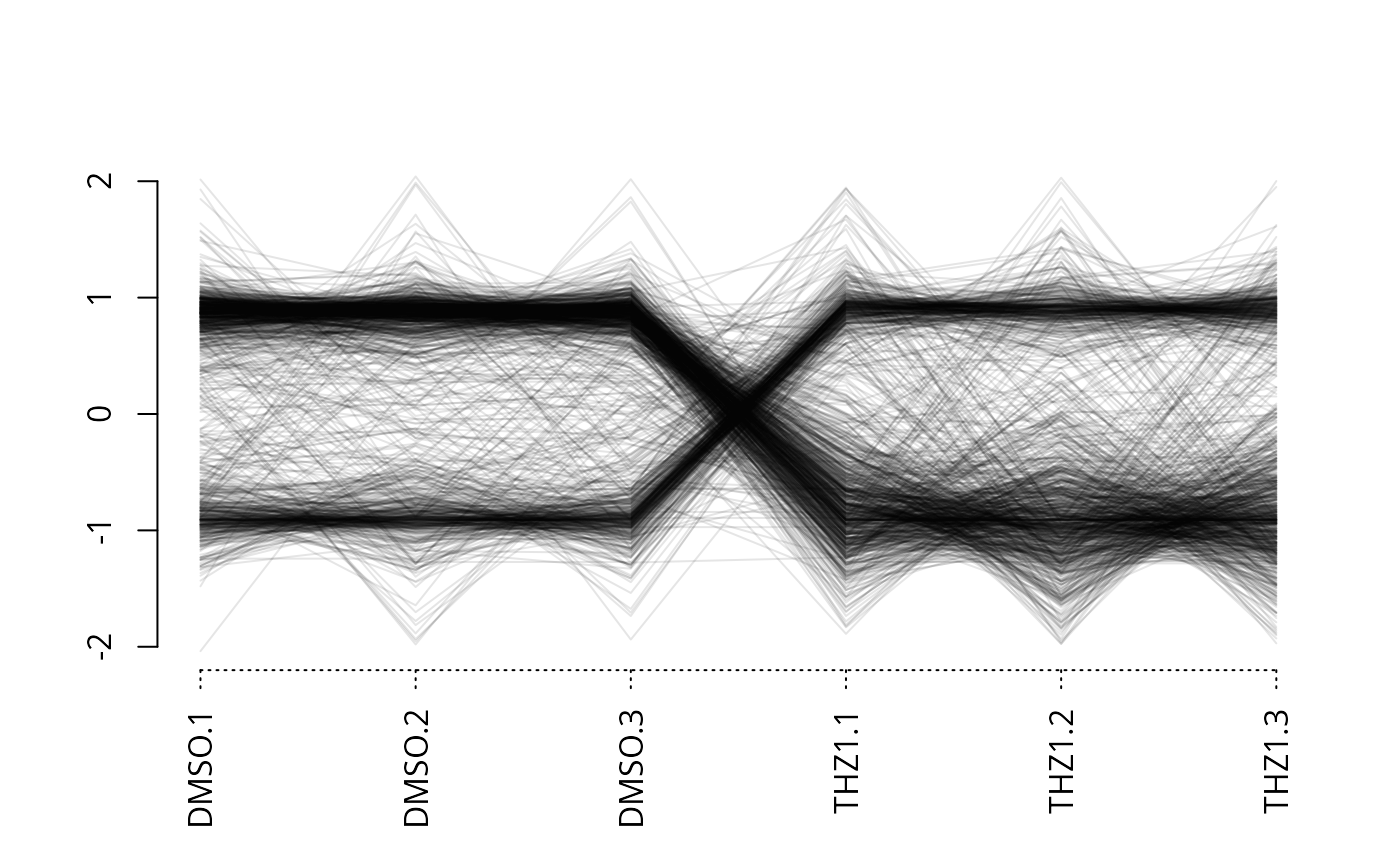
Parallel coordinates plot of row data for each column in a matrix
Source:R/plot-parallel2.R
plot_parallel2.RdCreate a parallel coordinates (scaled expression data on y-axis, samples on x-axis) for a matrix of data.
Usage
plot_parallel2(
x,
remove_var = 0.9,
scale = TRUE,
color = "black",
alpha = 0.1,
plot_title = NULL,
x_label = NULL,
y_label = NULL,
...
)Arguments
- x
feature x sample matrix
- remove_var
numeric. What proportion of low variance features to remove from the matrix before plotting. Default 0.9
- scale
bool. Should the data be standardized prior to plotting. Default TRUE
- color
character. Color of the lines. Default "black"
- alpha
numeric. Alpha value used to add transparency to lines. Default 0.1
- plot_title
character. title of the plot. Default NULL
- x_label
character. x-axis label
- y_label
character. y-axis label